AMD FidelityFX™ Single Pass Downsampler (SPD)
AMD FidelityFX Single Pass Downsampler (SPD) provides an AMD RDNA™ architecture optimized solution for generating up to 12 MIP levels of a texture.
Our latest RDNA™ 3 GPUs provide the ability to accelerate Generalized Matrix Multiplication (GEMM) operations. This means that you can now get hardware-accelerated matrix multiplications that take maximum advantage of our new RDNA 3 architecture. This new feature is called Wave Matrix Multiply Accumulate (WMMA).
This blog is a quick how-to guide for using the WMMA feature with our RDNA 3 GPU architecture using a Hello World example. It shows how to use the WMMA as a compiler intrinsic in HIP. As a prerequisite, we recommend reading through table 2 in section 1.1 of the RDNA 3 ISA guide for an overview of the various terminologies used. It is also recommended to go through the details of how WMMA works in the sections below before jumping straight to the source code examples. As a supplement to this blog, you can also refer to the AMD Matrix Instruction Calculator tool to generate in-depth information such as register mappings for every WMMA instruction available.
AMD GPUs based on the RDNA 3 architecture execute WMMA instructions in a very efficient manner allowing applications to achieve excellent performance and utilization. A single WMMA instruction coordinates 32 clocks of optimal work scheduling. AMD’s Mike Mantor, Corporate Fellow and Chief GPU Architect explains it like this:
The WMMA instruction optimizes the scheduling of data movement and peak math operations with minimal VGPR access by providing source data reuse and intermediate destination data forwarding operations without interruption. The regular patterns experienced in matrix operations enable WMMA instructions to reduce the required power while providing optimal operations that enable sustained operations at or very near peak rates.
WMMA supports inputs of FP16 or BF16 that can be useful for training online or offline, as well as 8-bit and 4-bit integer data types suitable for inference. The table below compares the theoretical FLOPS/clock/CU (floating point operations per clock, per compute unit) of our flagship Radeon RX 7900 XTX GPU based on the RDNA 3 architecture over the previous flagship Radeon RX 6950 XT GPU based on RDNA 2 for different data types:
(IU8 and IU4 refers to the unsigned 8-bit integer datatype and unsigned 4-bit integer datatype respectively)
| Data type | RX 6950 XT FLOPS/clock/CU | RX 7900 XTX FLOPS/clock/CU |
|---|---|---|
| FP16 | 256 | 512 |
| BF16 | N/A | 512 |
| IU8 | 512 | 512 |
| IU4 | 1024 | 1024 |
Unlike traditional per-thread matrix multiplication, WMMA allows the GPU to perform matrix multiplication cooperatively across an entire wavefront of 32 threads in wave32 mode or 64 threads in wave64 mode. This provides the benefit of sharing input/output matrix data across lanes of a wave, thus optimizing the VGPR usage and reducing memory traffic.
Suppose we have the GEMM operation using matrices A, B, C, and D:
D = A*B + C
where A and B are the input matrices, C is the accumulator matrix, and D is the destination matrix, also known as the result matrix.
If C isn’t used (e.g. in cases where you don’t use biases in your neural network), you can initialize C to 0 and re-use it as the result matrix:
C = A*B + C
This can be illustrated in the figure below, where matrices A, B, C, and D are all using a tile size of 16x16:
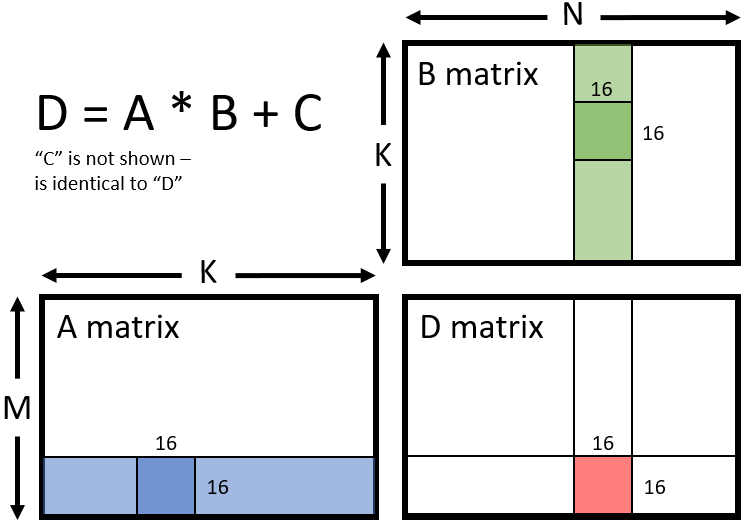
As of writing this blog, the three ways you can use WMMA can be via the compiler intrinsic which is available in LLVM clang built-ins, or writing inline assembly on your own, or you can also use rocWMMA, which will allow developers to get access to WMMA-based matrix operations (more details towards the end of this blog). We will focus on the compiler intrinsic approach in this blog.
The WMMA compiler intrinsic follows a certain syntax which is described follows:
D_frag = __builtin_amdgcn_wmma_<C, D format>_16x16x16_<A, B format>_w<32 or 64>(A_frag, B_frag, C_frag, OPSEL)
If you want to re-use D as C, where C is initialized to zero, simply replace D with C:
C_frag = __builtin_amdgcn_wmma_<C, D format>_16x16x16_<A, B format>_w<32 or 64>(A_frag, B_frag, C_frag, OPSEL)
Here, the “C, D format” refers to the format of matrices C and D respectively, which can be any one of f16, f32, or bf16 for floating point datatypes, and i32 as an integer datatype.
The “A,B format” refers to the input matrices A and B respectively, the format of which can be any one of f16, bf16, iu8, or iu4.
The 16x16x16 represents the GEMM convention for the tile size for a MxNxK matrix multiply, where matrix A is of size MxK, matrix B of size KxN, and matrix C/D is of size MxN. In the case of RDNA 3, only 16x16 tile sizes are supported. If your matrix is larger than 16x16, then split it into chunks of 16x16 which can then be passed into the WMMA instruction. In the context of a wave, internally the WMMA instruction takes a tile of 16x16 for matrix A and a tile of 16x16 for matrix B. It then multiplies them to give a 16x16 tile, which is then added with matrix C to give the final 16x16 matrix D tile.
The w<32 or 64> in the intrinsic describes whether WMMA is running in wave32 mode or wave64 mode. Depending on the mode, the loading and storing behavior of the matrices may vary. We will describe the differences later in this blog.
The final parameter “OPSEL” will also be explained a little later in this blog. For now, let’s focus on how these matrix fragments (A_frag, B_frag, C_frag, and D_frag) are loaded and used.
The A_frag, B_frag, C_frag, and D_frag parameters are the matrix fragments holding 16 elements each of matrices A, B, C and D respectively. From the perspective of a single lane (thread) within a wave, each “fragment” is locally stored in VGPRs, with each VGPR being 32 bits wide. Each thread holds A_frag and B_frag in 8 VGPRs for fp16/bf16, 4 VGPRs for iu8, and 2 VGPRs for iu4 regardless of the wave size.
C_frag and D_frag requires 8 VGPRs in wave32 mode and 4 VGPRs in wave64 mode, regardless of the datatype used by matrices C and D.
It is important to note that WMMA on RDNA 3 requires that the contents of A_frag and B_frag are replicated between lanes 0-15 and lanes 16-31 of the wave in wave32 mode. This means that for wave32 mode, each VGPR in lane 0 must have the exact same matrix data as each VGPR in lane 16. It is similar for lane 1 into lane 17, and so on, all the way to lane 15 into lane 31. This effectively maintains two copies of matrix data between the two half-waves. In wave64 mode, data from lanes 0-15 must also be replicated into lanes 32-47 and 48-63.
There are currently 12 such WMMA intrinsics following the above syntax. They are broadly divided into two categories, wave32 and wave64, described below:
| wave32 | wave64 | Matrix A,B format | Matrix C,D format |
|---|---|---|---|
__builtin_amdgcn_wmma_f32_16x16x16_f16_w32 | __builtin_amdgcn_wmma_f32_16x16x16_f16_w64 | FP16 | FP32 |
__builtin_amdgcn_wmma_f32_16x16x16_bf16_w32 | __builtin_amdgcn_wmma_f32_16x16x16_bf16_w64 | BF16 | FP32 |
__builtin_amdgcn_wmma_f16_16x16x16_f16_w32 | __builtin_amdgcn_wmma_f16_16x16x16_f16_w64 | FP16 | FP16 |
__builtin_amdgcn_wmma_bf16_16x16x16_bf16_w32 | __builtin_amdgcn_wmma_bf16_16x16x16_bf16_w64 | BF16 | BF16 |
__builtin_amdgcn_wmma_i32_16x16x16_iu8_w32 | __builtin_amdgcn_wmma_i32_16x16x16_iu8_w64 | IU8 | I32 |
__builtin_amdgcn_wmma_i32_16x16x16_iu4_w32 | __builtin_amdgcn_wmma_i32_16x16x16_iu4_w64 | IU4 | I32 |
Finally, the “OPSEL” parameter is a boolean flag required to be specified when using a 16-bit format for the C and D matrices. If this flag is set to true, the elements of C and D are stored in the upper half of the VGPR. However, when this flag is set to false, they are stored in the lower half of the VGPR. If you prefer 0-indexing, set this flag to false. This is illustrated in the code snippet taken from our Hello World example below, where we are storing 16-bit elements from C_frag into matrix C:
OPSEL pseudocode
// call the WMMA intrinsic with OPSEL set to "false"c_frag = __builtin_amdgcn_wmma_f16_16x16x16_f16_w32(a_frag, b_frag, c_frag, false);// 8 VGPRs per C,D fragment per thread in wave32 modeconst int lane = threadIdx.x % 16;
for (int ele = 0; ele < 8; ++ele){ // index into matrix C const int r = ele * 2 + (lIdx / 16); // store results from unpacked c_frag output c[16 * r + lane] = c_frag[ele*2];}Note the line that stores the matrix C elements from C_frag when OPSEL is set to “false”:
OPSEL=“false”
c[16 * r + lane] = c_frag[ele*2];If OPSEL was set to “true”, then the line above would instead be:
OPSEL=“true”
c[16 * r + lane] = c_frag[ele*2 + 1];Note that in this particular example, we choose matrix C to always store the C_frag elements in packed form, so the OPSEL flag should only affect the indexing in the right-hand side of this expression. You are free to modify this to store in unpacked format based on your app’s requirements.
WMMA requires a combination of row-major and column-major inputs for matrices A, B, C, and D. Matrix A is stored in column-major order, whereas matrices B, C, and D are all stored in row-major order. Matrices A and B are stored in packed format (i.e. each VGPR packs 2 fp16 values, 4 iu8 values, or 8 iu4 values), whereas matrices C and D are stored in unpacked format, with the “OPSEL” parameter used to describe the location of storage within the VGPR, described further below.
w32 or w64 represents wave32 or wave64 respectively. It represents the number of threads that will participate in the 16x16x16 GEMM operation.
Here we will demonstrate how to use __builtin_amdgcn_wmma_f16_16x16x16_f16_w32 to perform a 16x16x16 GEMM with fp16 inputs and outputs in wave32 mode.
The following figure shows the input matrix layout for matrices A and B. For matrix A, each (i, j) in a cell represents the i-th row and j-th column. For matrix B, each (i, j) in a cell represents the i-th column and j-th row.
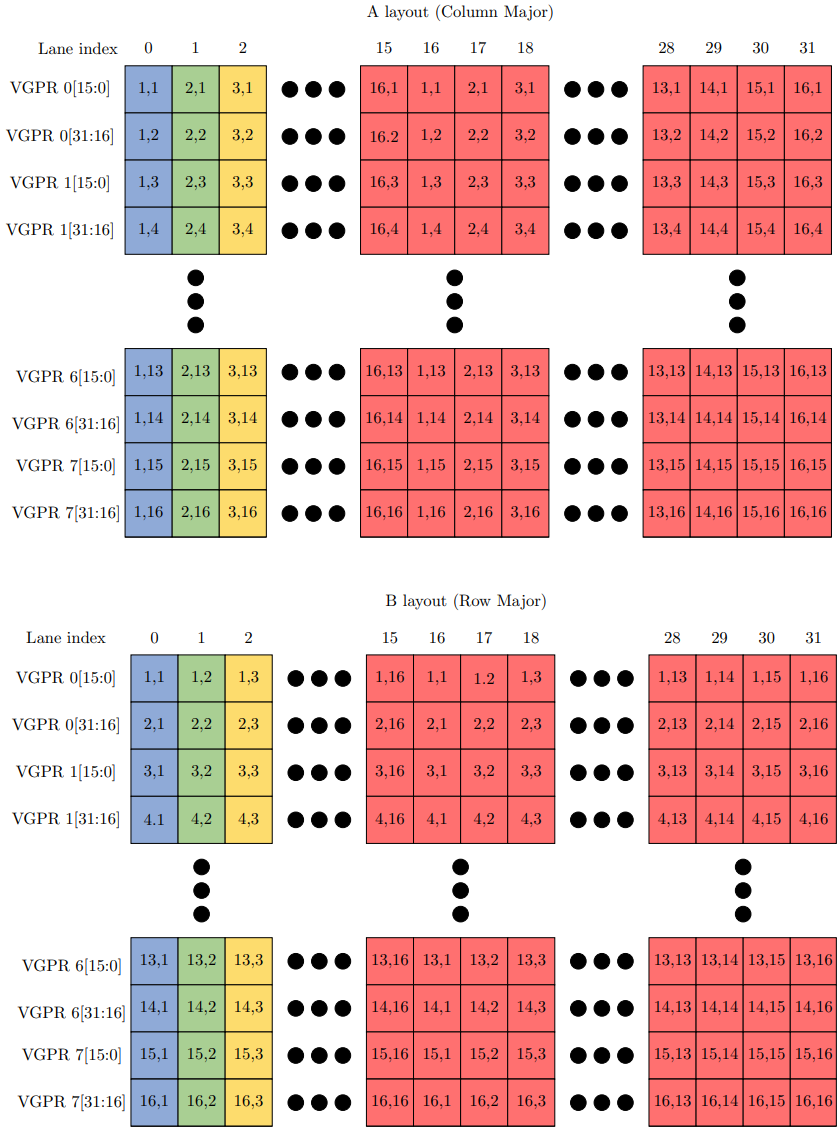
From the perspective of a thread, each VGPR holds two packed fp16 elements, with each set of 8 VGPRs holding 16 elements for matrices A and B respectively. Matrix A holds 16 columns in VGPRs whereas matrix B holds 16 rows in VGPRs.
Note the blue, green, and yellow colored cells representing 3 rows of the matrix A and similarly 3 columns of matrix B. These will be mapped to the following figure which shows the layouts of matrices C and D. Also note here the 8 VGPRs per lane would store the elements of C and D in unpacked format, with the 16 bit elements stored in the upper or lower half of the 32-bit VGPR based on the “OPSEL” flag. In our case, OPSEL is set to 0 (False) so each VGPR holds a matrix element in the lower half of the VGPR (bits 0 to 15). Note that, as mentioned before, matrices C and D here are stored in row-major format.
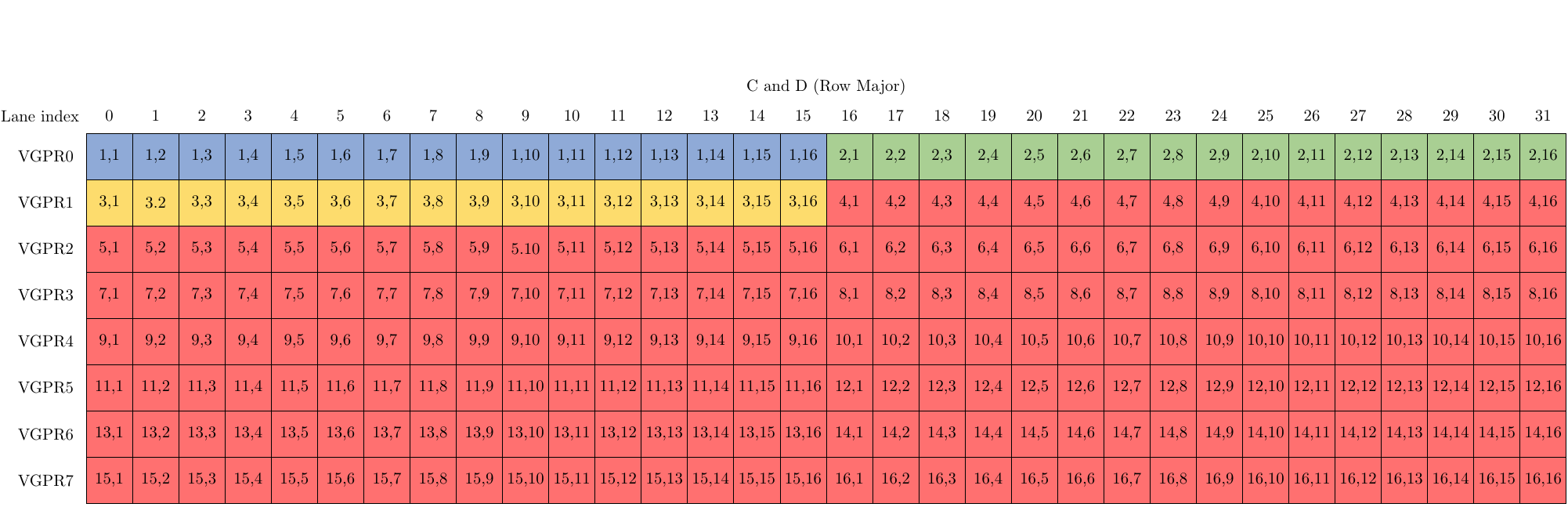
The following is a code example with some helpful comments, showing how to perform a matrix multiplication of two f16 matrices A and B, and re-using C as D for the GEMM operation C = AB + C in wave32 mode using __builtin_amdgcn_wmma_f16_16x16x16_f16_w32
wmma_test.cpp:
WMMA example
// Wave Matrix Multiply Accumulate (WMMA) using HIP compiler intrinsic// Does a matrix multiplication of two 16x16, fp16 matrices, and stores them into a 16x16 fp16 result matrix
#include <iostream>#include <hip/hip_runtime.h>#include <hip/hip_fp16.h>
using namespace std;
// Use half16 as an alias of the internal clang vector type of 16 fp16 valuestypedef _Float16 half16 __attribute__((ext_vector_type(16)));
__global__ void wmma_matmul(__half* a, __half* b, __half* c){ const int gIdx = blockIdx.x * blockDim.x + threadIdx.x; const int lIdx = threadIdx.x;
// a and b fragments are stored in 8 VGPRs each, in packed format, so 16 elements each for a and b // a_frag will store one column of the 16x16 matrix A tile // b_frag will store one row of the 16x16 matrix B tile half16 a_frag; half16 b_frag; // initialize c fragment to 0 half16 c_frag = {};
// lane is (0-31) mod 16 instead of 0-31 due to matrix replication in RDNA 3 const int lane = lIdx % 16;
for (int ele = 0; ele < 16; ++ele) { b_frag[ele] = b[16*ele + lane]; }
for (int ele = 0; ele < 16; ++ele) { a_frag[ele] = a[16 * lane + ele]; }
// call the WMMA intrinsic with OPSEL set to "false" c_frag = __builtin_amdgcn_wmma_f16_16x16x16_f16_w32(a_frag, b_frag, c_frag, false);
for (int ele = 0; ele < 8; ++ele) { const int r = ele * 2 + (lIdx / 16); // store results from unpacked c_frag output c[16 * r + lane] = c_frag[ele*2]; // if OPSEL was set to "true", the line above would instead be // c[16 * r + lane] = c_frag[ele*2 + 1]; }
}
int main(int argc, char* argv[])
{ __half a[16 * 16] = {}; __half b[16 * 16] = {}; __half c[16 * 16] = {}; __half *a_gpu, *b_gpu, *c_gpu; hipMalloc(&a_gpu, 16*16 * sizeof(__half)); hipMalloc(&b_gpu, 16*16 * sizeof(__half)); hipMalloc(&c_gpu, 16*16 * sizeof(__half));
// fill in some data into matrices A and B for (int i = 0; i < 16; ++i) { for (int j = 0; j < 16; ++j) { a[i * 16 + j] = (__half)1.f; b[i * 16 + j] = (__half)1.f; } }
hipMemcpy(a_gpu, a, (16*16) * sizeof(__half), hipMemcpyHostToDevice); hipMemcpy(b_gpu, b, (16*16) * sizeof(__half), hipMemcpyHostToDevice); hipMemcpy(c_gpu, c, (16*16) * sizeof(__half), hipMemcpyHostToDevice);
wmma_matmul<<<dim3(1), dim3(32, 1, 1), 0, 0>>>(a_gpu, b_gpu, c_gpu);
hipMemcpy(c, c_gpu, (16 * 16) * sizeof(__half), hipMemcpyDeviceToHost);
hipFree(a_gpu); hipFree(b_gpu); hipFree(c_gpu);
for (int i = 0; i < 16; ++i) { for (int j = 0; j < 16; ++j) { printf("%f ", (float)c[i * 16 + j]); } printf("\\n"); }
return 0;}The process in the above code is as follows:
Initialize the input matrices A and B.
Set matrix C to zero and re-use it as matrix D.
Pass matrix C to the “wmma_matmul” kernel, which loads the matrix elements into their respective fragments.
Call the WMMA intrinsic.
Store the result from c_frag into matrix C.
To compile the above program on your Radeon RX 7900 XTX or 7900 XT GPU on Linux or Windows using HIP, simply use hipcc --offload-arch=gfx1100 wmma_test.cpp -o wmma_test. Make sure you have ROCm v5.4 or newer installed on your Linux environment, or the latest HIP SDK installed on your Windows® environment.
As an alternative to installing the HIP SDK, head to the Orochi GitHub repository for an example involving usage of hipRTC APIs to compile and run the above code at runtime on Windows® or Linux!
On a side note, if you’re used to using nvcuda::wmma APIs and/or rocWMMA, you will notice many similarities here. For example, these matrix fragments in a_frag, b_frag, c_frag, and d_frag can be considered the same as the fragment templated type available in those APIs, with the loading and storing of fragments similar to load_matrix_sync and store_matrix_sync respectively. The call to the compiler intrinsic is similar to calling mma_sync. The main difference here is that you are doing the loading, storing, syncing, and WMMA calls yourself, rather than relying on the API to do it for you. For brevity, we’ve skipped the synchronization part as it is not needed for a simple example such as the one above, however we do recommend using __syncthreads() wherever appropriate.
WMMA can be used to accelerate any use case that involves matrix multiplication. Here we describe three such use cases that are either already available or will be coming soon:
Stable diffusion uses WMMA to boost its performance via the SHARK MLIR/IREE runtime for RDNA 3 GPUs
AMD’s Composable Kernels (CK) library will soon be updated in a new release to support WMMA, which will enable Meta’s AI Template (AIT) library to support end-to-end hardware acceleration for model inference on RDNA 3.
The Machine Intelligence Shader Autogen (MISA) library will soon release WMMA support to accelerate models like Resnet50 for a performance uplift of roughly 2x from RDNA 2.
So far, we’ve discussed how to use WMMA via compiler intrinsics. However, it may be cumbersome to integrate this with existing CUDA-based applications that utilize the nvcuda::wmma APIs via. mma.h (note that the WMMA in nvcuda::wmma refers to Warp Matrix-Multiply Accumulate, which is different from the Wave Matrix-Multiply Accumulate described here).
While it’s true that these intrinsics can be mapped easily to the mma_sync API call, the matrix loading/storing and synchronization can be tricky to handle and debug, especially for novice users.
RDNA 3 WMMA support is now available in rocWMMA. This library is portable with nvcuda::wmma and it supports MFMA and WMMA instructions, thus allowing your application to have hardware-accelerated ML in both RDNA 3 and CDNA 1/2 based systems.
Thanks to Atsushi Yoshimura, Joseph Greathouse and Chris Millette for suggesting improvements and providing feedback, and thanks to Mike Mantor for contributing the insightful WMMA explanations. Matrix layout figures generated by using modified TikZ programs originally written by Damon McDougall.

